- Press Office
- Press releases 2021
- How ethane-consuming microbes pick up their favorite dish
How ethane-consuming microbes pick up their favorite dish
This insight is the result of the close collaboration of several research groups at the Max Planck Institute for Marine Microbiology. A team around the PhD student Cedric Hahn and Gunter Wegener recently discovered ethane-degrading microbes at hydrothermal vents of the Guaymas Basin at a water depth of 2,000 meters in the Gulf of California. They named it Ethanoperedens thermophilum, which means “heat-loving ethane-eater”. Cedric Hahn cultured the ethane-degrading microbes in the laboratory.
Hahn and Wegener colleagues of the research group Microbial Metabolism, Tristan Wagner and Olivier Lemaire took a closer look at these microorganisms. This collaborative work unraveled the secrets behind the ethane fixation. “We were astonished by what we found. Besides a global similarity, some features of the enzyme differ fundamentally from its counterpart – the enzyme responsible for the degradation of methane,” says Gunter Wegener, scientist in the Research Group Deep-Sea Ecology and Technology.
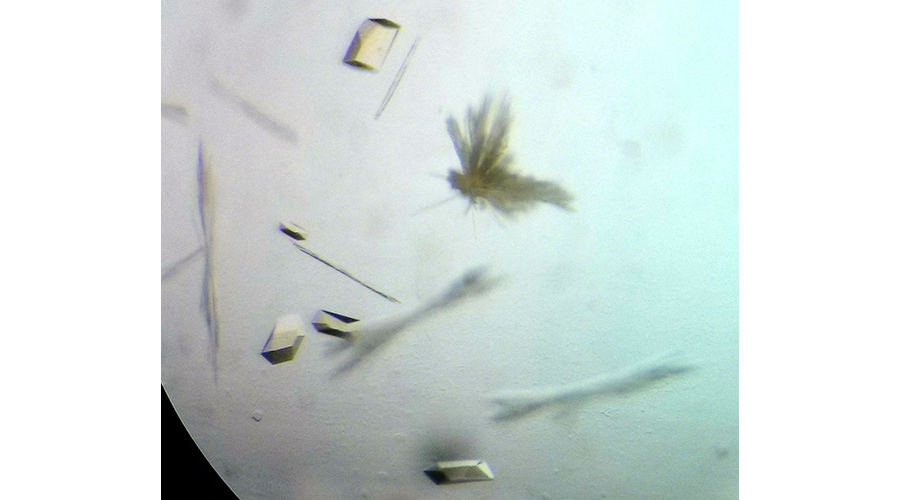

(©Max-Planck-Institut für Marine Mikrobiologie, O. Lemaire und T. Wagner)
Ethane-eaters depend on the same enzyme as Methane-eaters
In deep-sea sediments, geothermal heat leads to the degradation of organic matter to oil and natural gases such as ethane. The ethane is consumed by different microorganisms that form a so-called consortium: archaea, which break down the natural gas, and bacteria, which couple the electrons released in the process to the reduction of sulfate, an abundant compound in the ocean. The discovery of the ethane-eating microbes has brought a breath of fresh air to the research. Compared to microbes eating methane that take a lot of time to grow, the ethane specialists grow much faster and double every week. Thus, the time of biomass production is reduced, which allow attempts of purification and characterization of key enzymes catalyzing the oxidation of natural gas.
To test similarities between the enzymes catalyzing the activation of ethane and methane, Cedric Hahn added a well-known molecular inhibitor of methane oxidation to his culture. This treatment abolished also ethane oxidation. “This suggested that the ethane-oxidizing archaea active ethane in similar enzymatic reactions as those acting in methane degradation/generation” says Cedric Hahn. Such enzymes are a key expertise of Tristan Wagner who studied them for several years.
A structure visualized at an astonishing level of precision
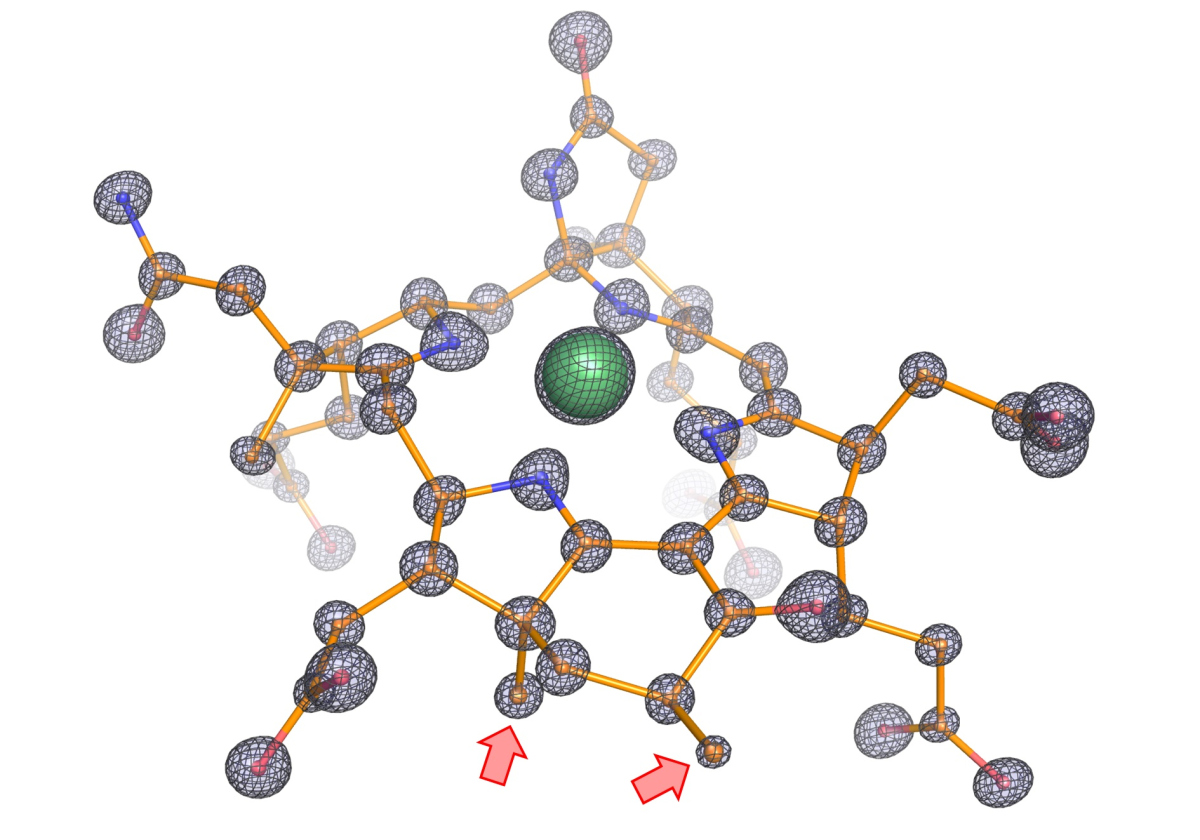
(©Max-Planck-Institut für Marine Mikrobiologie, O. Lemaire und T. Wagner)
Cedric Hahn and Olivier Lemaire, the two first authors of the paper now published in Science, then tried to purify the enzyme responsible of ethane fixation. “The project was highly challenging,” says Olivier Lemaire. “Usually we purify the enzymes from much larger amounts of biomass from a culture containing one microorganism. However, we finally yielded sufficient amounts of pure enzymes for structural analyses.”
The next critical step was to obtain crystals of the enzyme to determine its tri-dimensional structure. “X-ray crystallography gave previous excellent results on this group of enzymes” says Tristan Wagner, head of the Microbial Metabolism research group and expert on this technique. “We analyzed these crystals by X-ray diffraction and solved the enzyme structure at an unprecedented atomic resolution. We can thus determine the position of individual atoms and obtain therefore an extremely precise picture of the structure”.
The structure exhibits several unprecedented features. “We noticed that the catalytic chamber in which the chemical reaction takes place is twice as large as in enzymes capturing methane, which makes sense considering that ethane is bigger than methane,” says Olivier Lemaire. The cofactor, catalyst of the reaction, contains two additional methyl groups. These methyl groups were confirmed by Jörg Kahnt from the Max Planck Institute for Terrestrial Microbiology, a worldwide expert on this cofactor. “We found a protein that could be responsible for these methylations, and it is only found in ethane-consumers,” says Cedric Hahn. Since the chamber is more voluminous, a normal cofactor would simply not fit properly and would impair the reaction. The methylations on the cofactor anchor it in the correct position.
Furthermore, the enzyme contains a tunnel connecting the exterior to the catalytic chamber. This tunnel does not exist in any characterized similar enzymes. The researchers experimentally proved the existence of this tunnel through a collaboration with Sylvain Engilberge at the Paul Scherrer Institute in Switzerland, where the protein crystals were gassed with xenon. Xenon was detected in the catalytic chamber and the predicted gas tunnel, proving its existence. The tunnel is maintained and stabilized by modified amino acids and additional extensions.
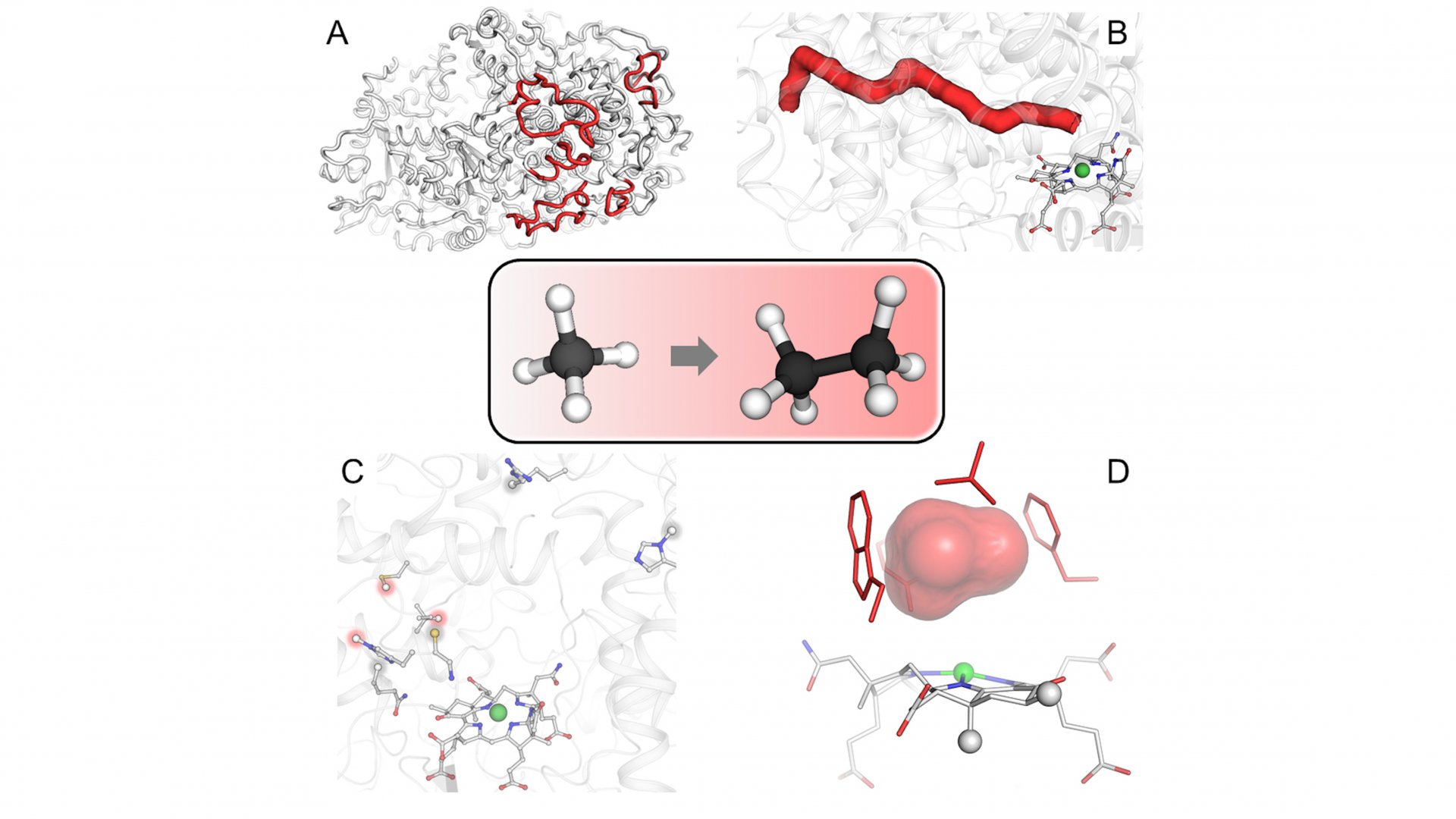
Now the spotlight turns to propane and butane
The enzyme structure illustrates how these microbes from geothermally active seeps became specialized in ethane capture. This work allows a deeper understanding of the first step in ethane degradation, the only source of energy of these archaea. “Our finding that the enzyme responsible for the process has specific traits to recognize ethane rather than other alkanes is a big step forward, however the understanding of the whole degradation process is still a long way off”, concludes Tristan Wagner.
So, how to proceed the research? “Our previous works show that the activation of longer alkanes requires similar enzymes” says Gunter Wegener. “As a next step, we want to investigate what could be the specific features of the enzymes that catalyze the activation of propane and butane”.
Original publication
Cedric J. Hahn*, Olivier N. Lemaire*, Jörg Kahnt, Sylvain Engilberge, Gunter Wegener, Tristan Wagner: Crystal structure of a key enzyme for anaerobic ethane activation, Science, July 2021
*Both authors contributed equally
Participating institutions
- Max Planck Institute for Marine Microbiology, Bremen, Germany
- Max Planck Institute for Terrestrial Microbiology, Marburg, Germany
- Paul Scherrer Institute (PSI), Villigen, Switzerland
- MARUM – Center for Marine Environmental Sciences, Bremen, Germany
Please direct your queries to:
Head of Group
MPI for Marine Microbiology
Celsiusstr. 1
D-28359 Bremen
Germany

Scientist
HGF MPG Joint Research Group for Deep-Sea Ecology and Technology
MPI for Marine Microbiology
Celsiusstr. 1
D-28359 Bremen
Germany
|
Room: |
1335 |
|
Phone: |
