- Departments
- Department of Molecular Ecology
Department of Molecular Ecology
Groups
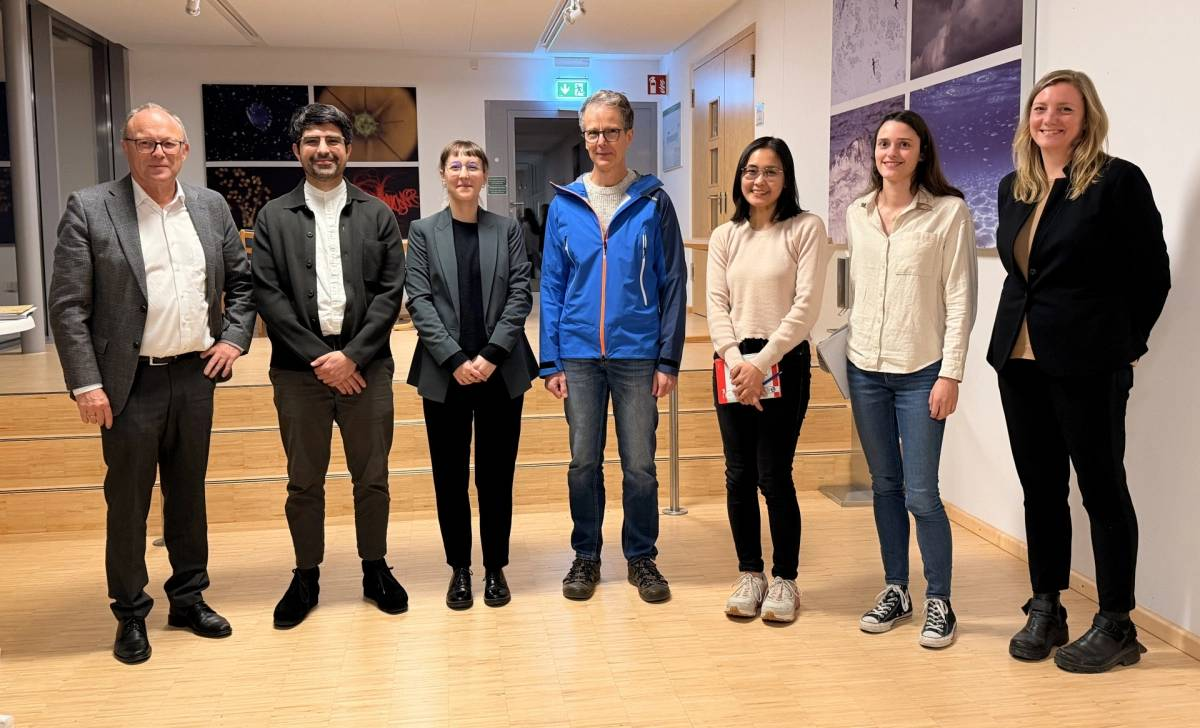
Inspiration and focus of our department
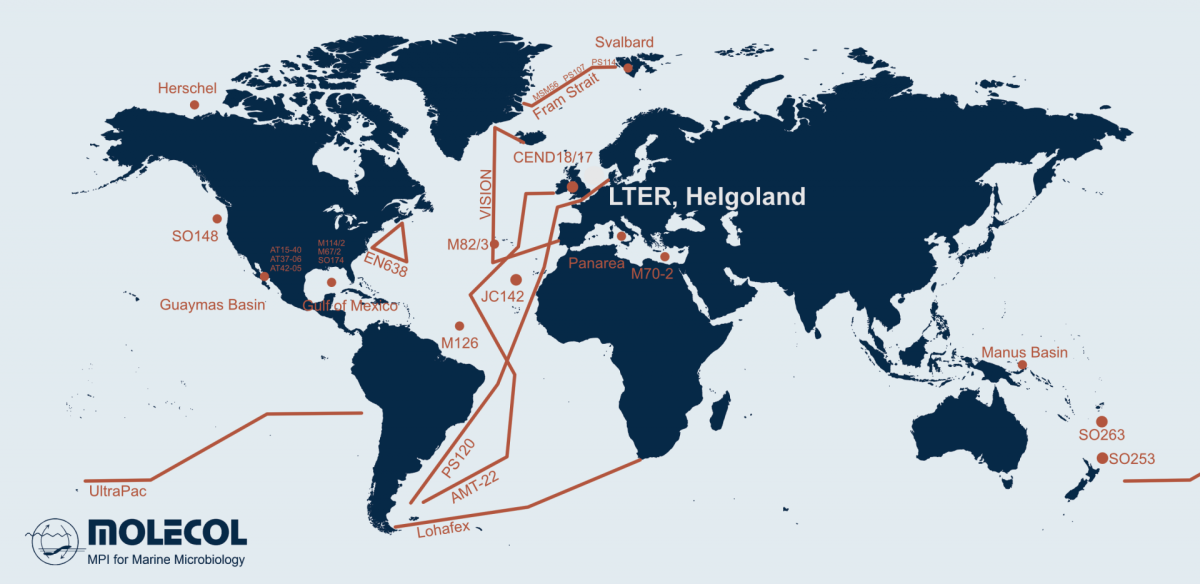
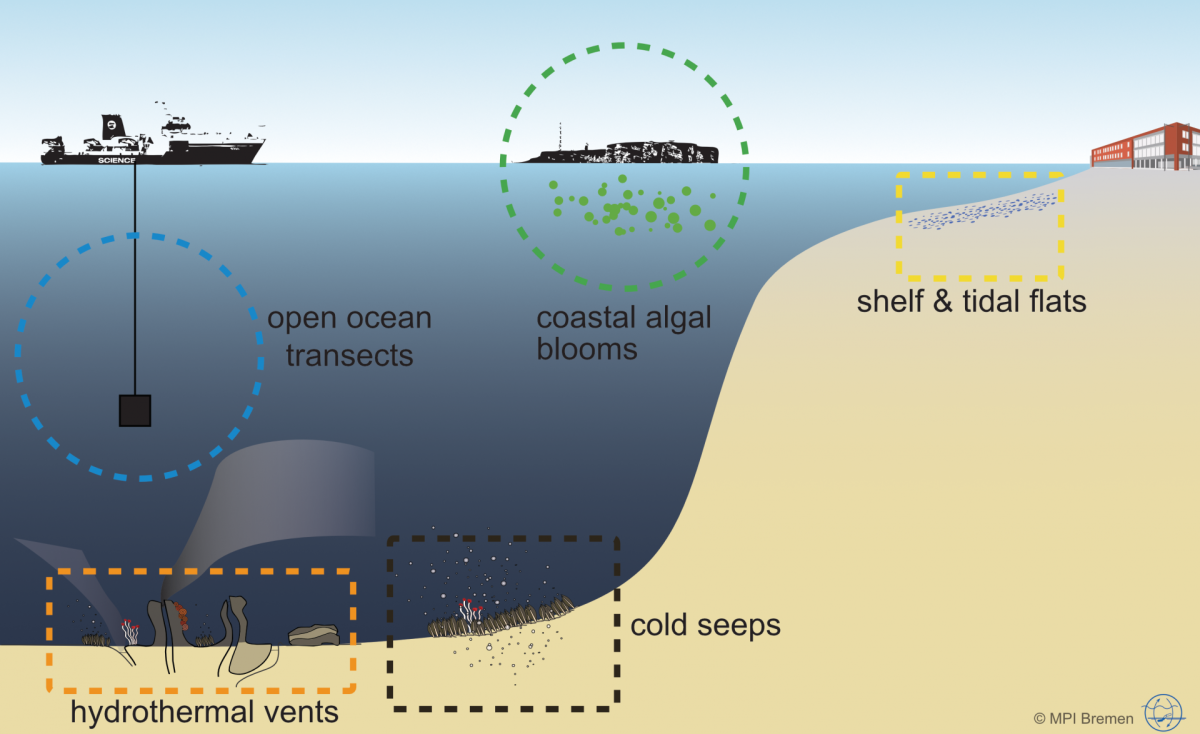
Microorganisms account for 70 % of the biomass in the oceans and are fundamental for the functioning of marine ecosystems. They play important roles in regulating organic matter and nutrient cycles and even influence the Earth's climate. Whereas only thousands of species of Bacteria and Archaea have been described by cultivation-based methods, we are identifying millions with molecular techniques. In our department, we use the most modern techniques in a multidisciplinary and polyphasic approach to ecologically characterise marine microbial communities from hydrothermal vents to coastal surface waters and from the tropics to the poles.
Projects
Managing Director
Department of Molecular Ecology
MPI for Marine Microbiology
Celsiusstr. 1
D-28359 Bremen
Germany
|
Room: |
2221 |
|
Phone: |

Assistance
Department of Molecular Ecology
MPI for Marine Microbiology
Celsiusstr. 1
D-28359 Bremen
Germany
|
Room: |
1345 |
|
Phone: |

News
"Congratulations to Isidora Morel on the successful defense of her doctoral dissertations"
[17.11.2025]
Uncultivated microbes in need of their own taxonomy
Taxonomy encompasses the identification, classification, and nomenclature of organisms. As such, taxonomy is a prerequisite for ecology. Only based on accurate taxonomic concepts and methods, can the diversity and composition of complex microbial communities be accurately described and monitored. Our department has a long tradition of developing and applying new taxonomic methods, from pioneering the in situ identification and quantification of not-yet cultivated bacteria and archaea through fluorescence in situ hybridization (FISH) to contributing to widely used standards of 16S rRNA gene classification.
More recently, the developments in sequencing technologies provide a vast amounts of information on yet uncultivated microorganisms. By combining multiple methods, such as metagenomics and FISH, we have been able to describe several new marine taxa including the species Candidatus Prosiliicoccus vernus (Francis et al. 2019) and the genus Candidatus Abditibacter (Grieb et al. 2020). To find out more about our work on the taxonomy of yet-uncultivated bacteria and archaea visit our project page for taxonomy.
Diversity, visualisation and cultivation
One of the primary questions of ecology is 'who or what is there?'. In marine microbial ecology, we typically employ high-throughput sequencing of the 16S rRNA gene to provide a window into the diversity of the microbial community. From this, we can determine the key microbial taxa present and begin to formulate hypotheses about ecological processes taking place. More recently, a significant advancement in long-read sequencing technology, with the advent of the PacBio Sequel II platform, now allows us to sequence the genetic material from environmental populations (metagenomics) and simultaneously retrieve full-length 16S rRNA genes. In comparison to short 16S rRNA gene amplicons, these full-length sequences can be used in more robust and accurate phylogenetic analyses.
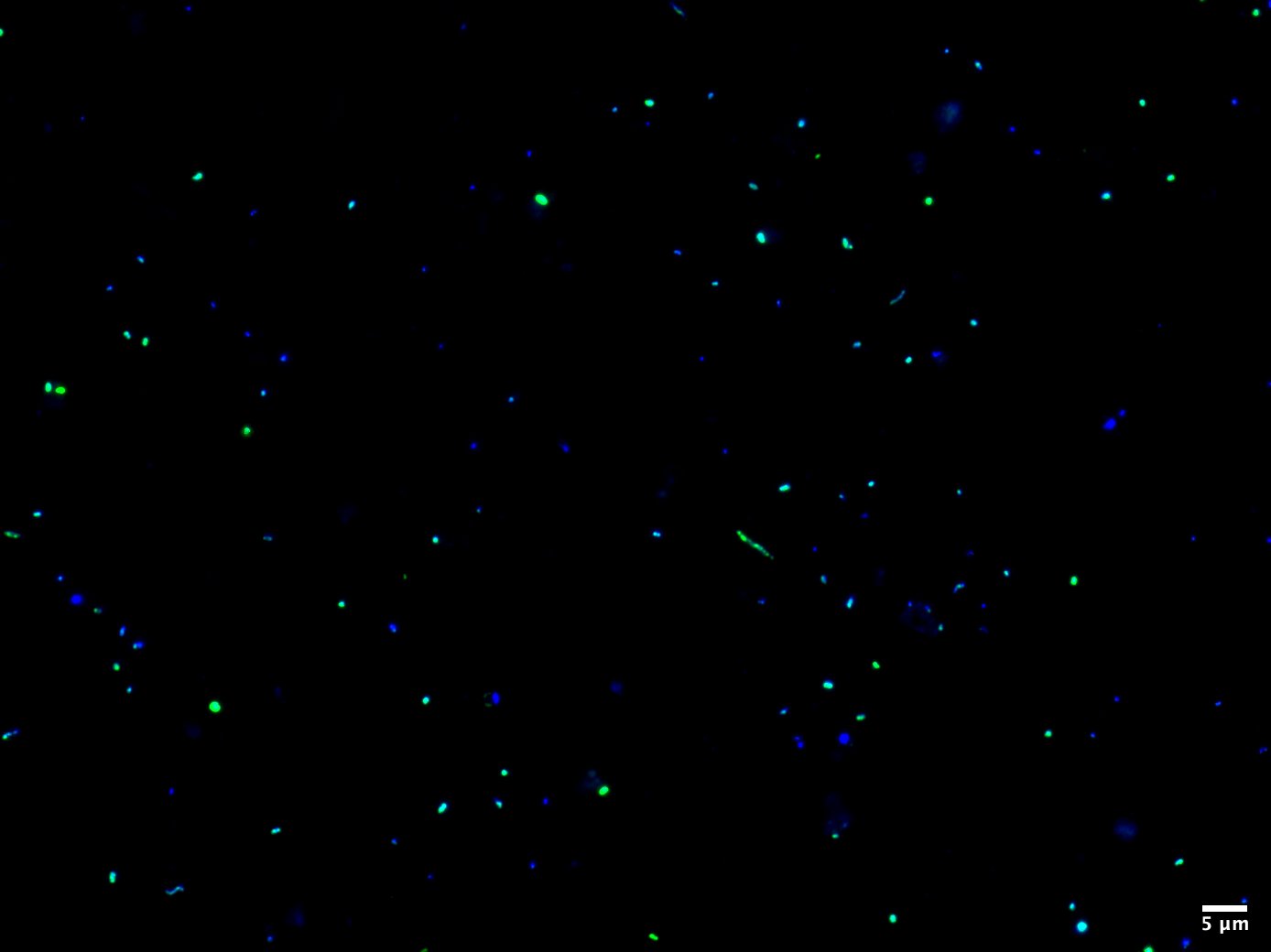
The next fundamental component of ecology is assessing the abundance of a specific population in the environment. Trivially speaking, the more individuals there are, the more important they are for the ecosystem and the higher the impact on the nutrient cycle, resource exploration and metabolic interactions. This is where our method FISH is an indispensable tool for the enumeration of taxonomically well-defined microorganisms. As most microbes lack a conspicuous morphology, the only reliable marker to distinguish between them is the ribosomal RNA molecule, which is highly conserved and abundant in every organism. Using small fluorescently labelled oligonucleotides complementary to the rRNA, we can tailor probes specific for the taxa of interest to visualize and count them by epifluorescence microscopy. Besides a numerical quantification, FISH provides a clue about cellular structures and physical interactions with other microbes. We can use high-resolution microscopy like Confocal Laser Scanning Microscopy (CLSM) and STimulated Emission Depletion (STED) microscopy to study intracellular properties or the uptake of, e.g., substrates. Using FISH and high-resolution microscopy, we can determine accurate cell-sizes and biovolumes of uncultivated microbes.
Though we can obtain ample amounts of information about the ecology of microorganisms from environmental samples, we ultimately still require pure cultures or enrichments in the lab. Cultivation is still, in most cases, the only way to experimentally verify predictions and hypotheses formulated from genomic sequence information. However, the isolation of marine microorganisms can be a challenging task, as it is difficult to recreate the natural environment and specific conditions required for a population to grow. Therefore, we use liquid mediums and enrichments along with physical separation of individual cells through dilution and molecular identification methods (in house developed specific PCR and in situ hybridisation probes) to isolate abundant microbes from natural communities. Once a strain is in pure culture, the source is unlimited and physiological traits and individual enzymes can be studied.

Functional characterization
The rapid development of massive sequencing technologies has unravelled a new era for the examination of the genetic potential of microbial communities. From the analyses of sequence discrete populations in nature to isolates in the lab, the use of multi-omic techniques allows us to integrate genetic information at the gene, transcript and also protein levels. Our goal is to bridge the gap between predicted function and measured microbial activity through the integration of these different levels of information.
The mRNA of species within microbial communities tells us which genes are actively transcribed whilst metaproteomics identifies the abundant proteins. Combining these observations enables us to predict the metabolic processes, only hampered by the fact that for half of the proteins we do not know their function. But the accumulated knowledge of mankind allows us a first glance at novel ecosystems. With the observation of community responses to substrate additions and the study of novel strains, we aim at a full understanding of the major processes in nature. We are still explorers.



